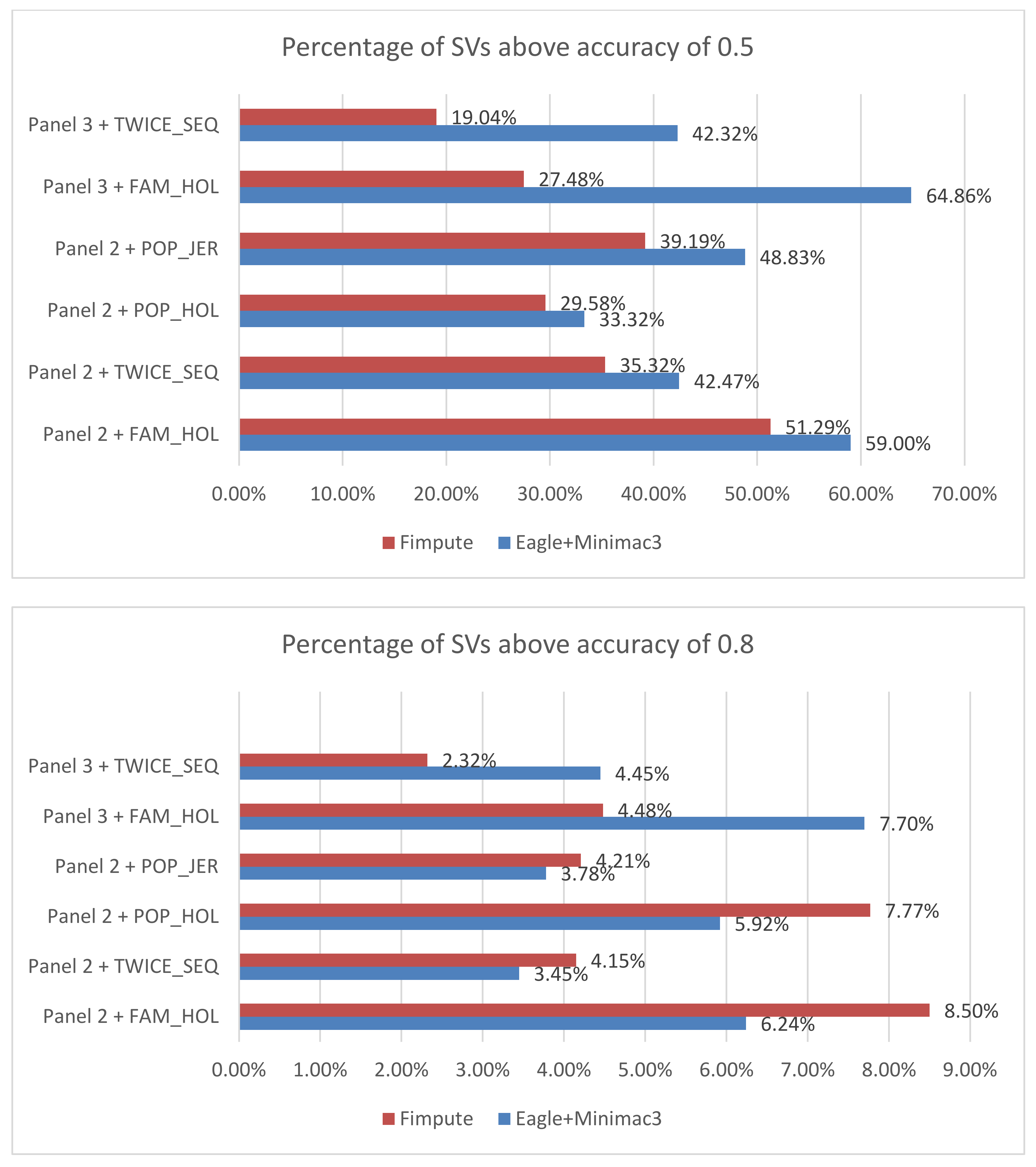
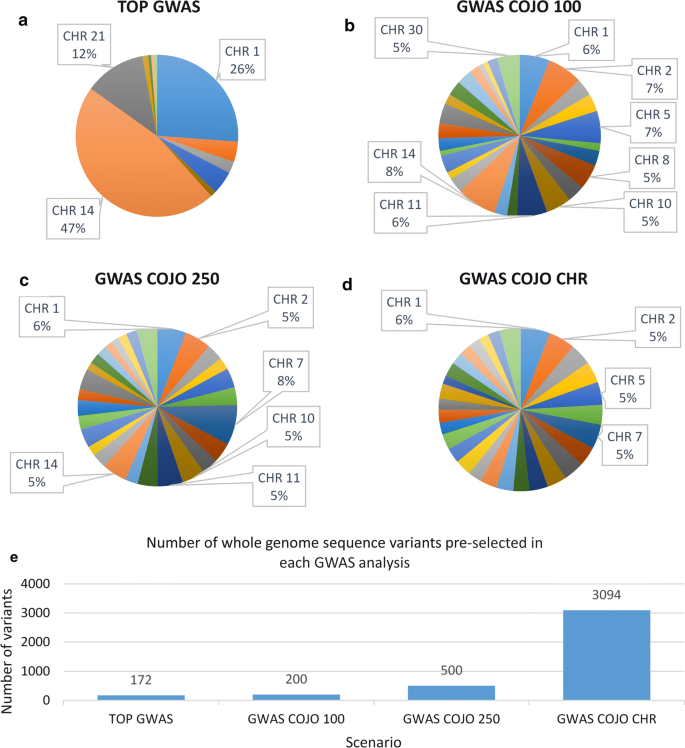
Animals | Free Full-Text | Investigating the Effect of Imputed Structural Variants from Whole-Genome Sequence on Genome-Wide Association and Genomic Prediction in Dairy Cattle | HTML
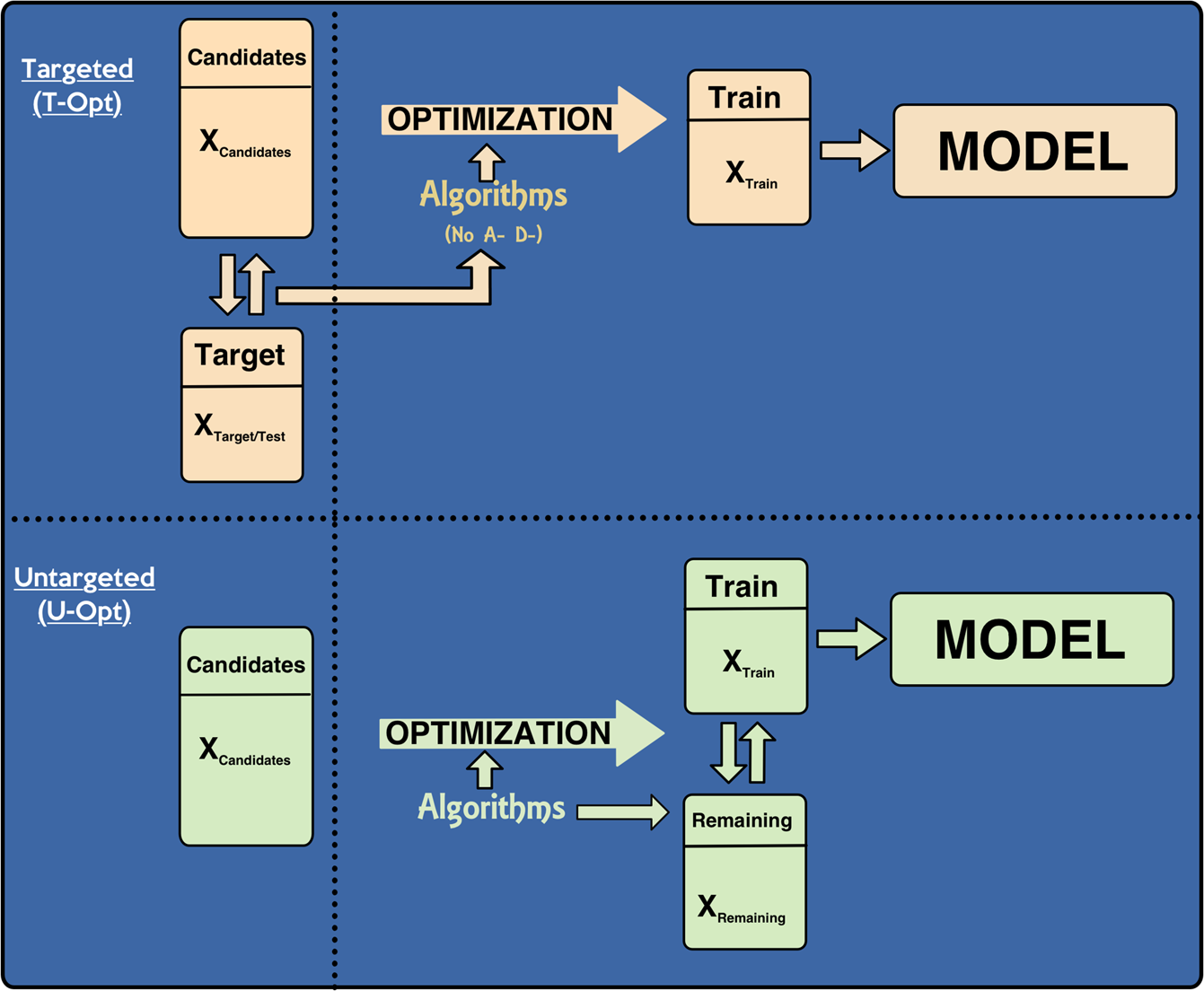
Use of whole-genome sequence data and novel genomic selection strategies to improve selection for age at puberty in tropically-adapted beef heifers | Genetics Selection Evolution | Full Text

Toward genomic prediction from whole-genome sequence data: impact of sequencing design on genotype imputation and accuracy of predictions | Heredity

Quantifying the contribution of sequence variants with regulatory and evolutionary significance to 34 bovine complex traits | PNAS

Added value of whole-genome sequence data to genomic predictions in dairy cattle Rianne van Binsbergen 1,2, Mario Calus 1, Chris Schrooten 3, Fred van. - ppt download
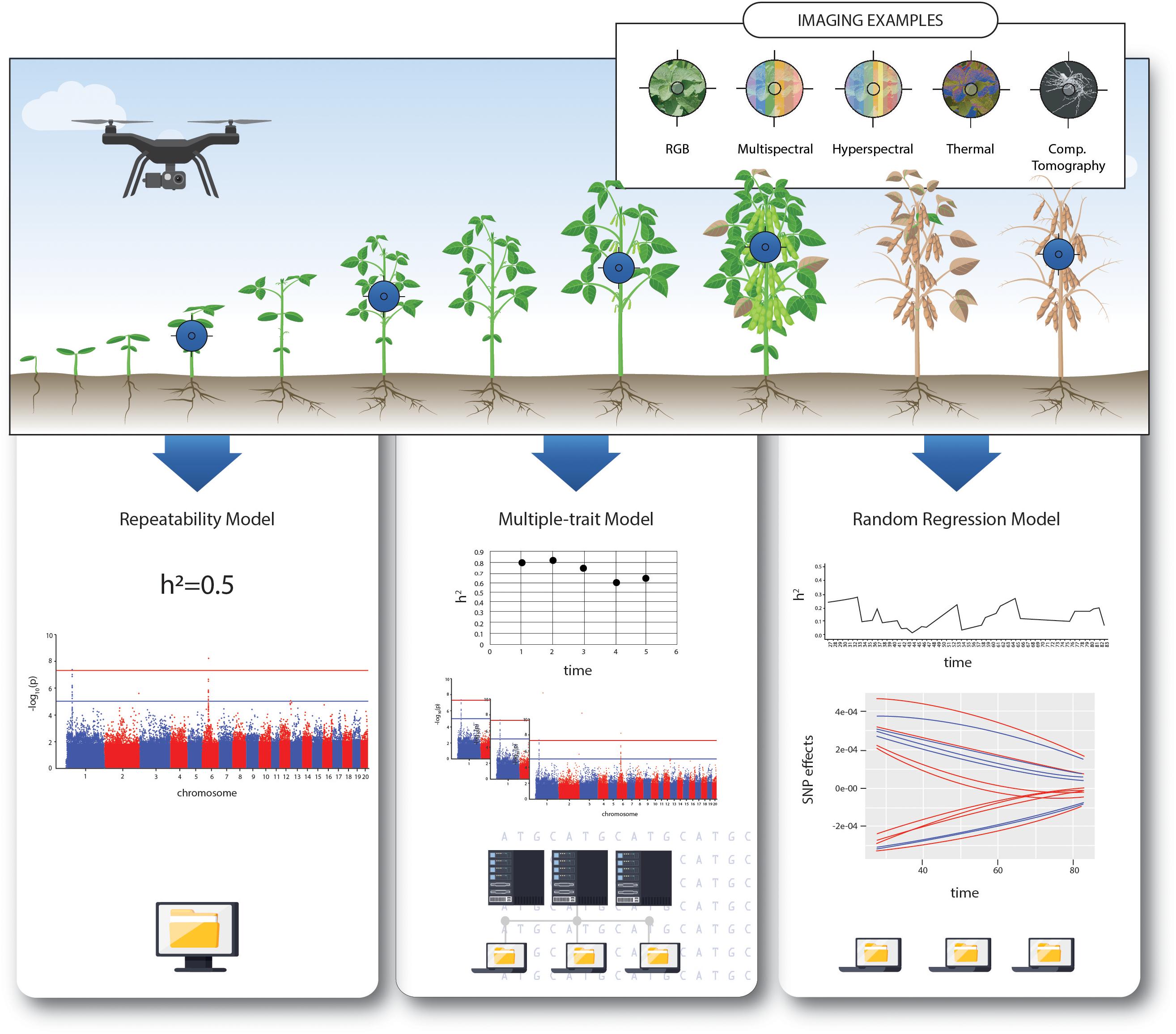
Frontiers | Integrating High-Throughput Phenotyping and Statistical Genomic Methods to Genetically Improve Longitudinal Traits in Crops | Plant Science

Exome sequencing highlights the role of wild-relative introgression in shaping the adaptive landscape of the wheat genome,Nature Genetics - X-MOL
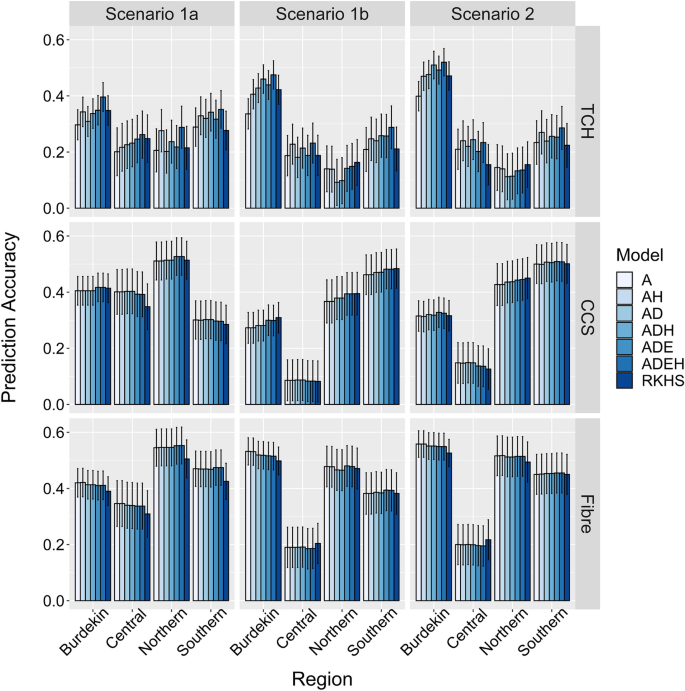
Improved genomic prediction of clonal performance in sugarcane by exploiting non-additive genetic effects | SpringerLink