PLOS Neglected Tropical Diseases: A comparison of three approaches for the discovery of novel tripartite attachment complex proteins in Trypanosoma brucei

The proximity-labeling technique BioID identifies sorting nexin 6 as a member of the insulin-like growth factor 1 (IGF1)–IGF1 receptor pathway - Journal of Biological Chemistry

Effective enrichment of surfaced displayed biotinylated AP. (a) Western... | Download Scientific Diagram
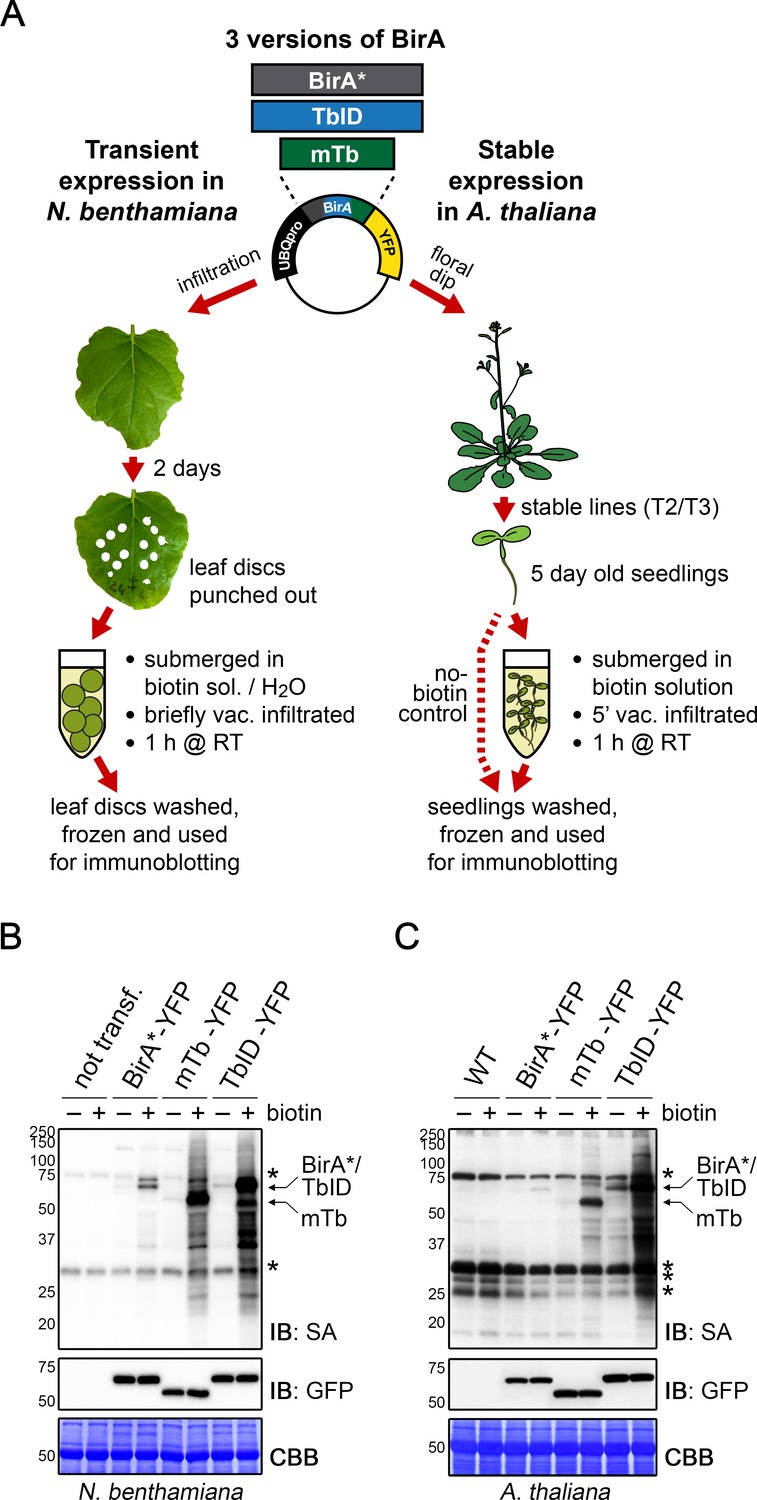
Proximity labeling of protein complexes and cell-type-specific organellar proteomes in Arabidopsis enabled by TurboID | eLife

A bacterial display system for effective selection of protein-biotin ligase BirA variants with novel peptide specificity | Scientific Reports

FLI-1 Functionally Interacts with PIASxα, a Member of the PIAS E3 SUMO Ligase Family* - Journal of Biological Chemistry

Mapping the micro-proteome of the nuclear lamina and lamina-associated domains | Life Science Alliance
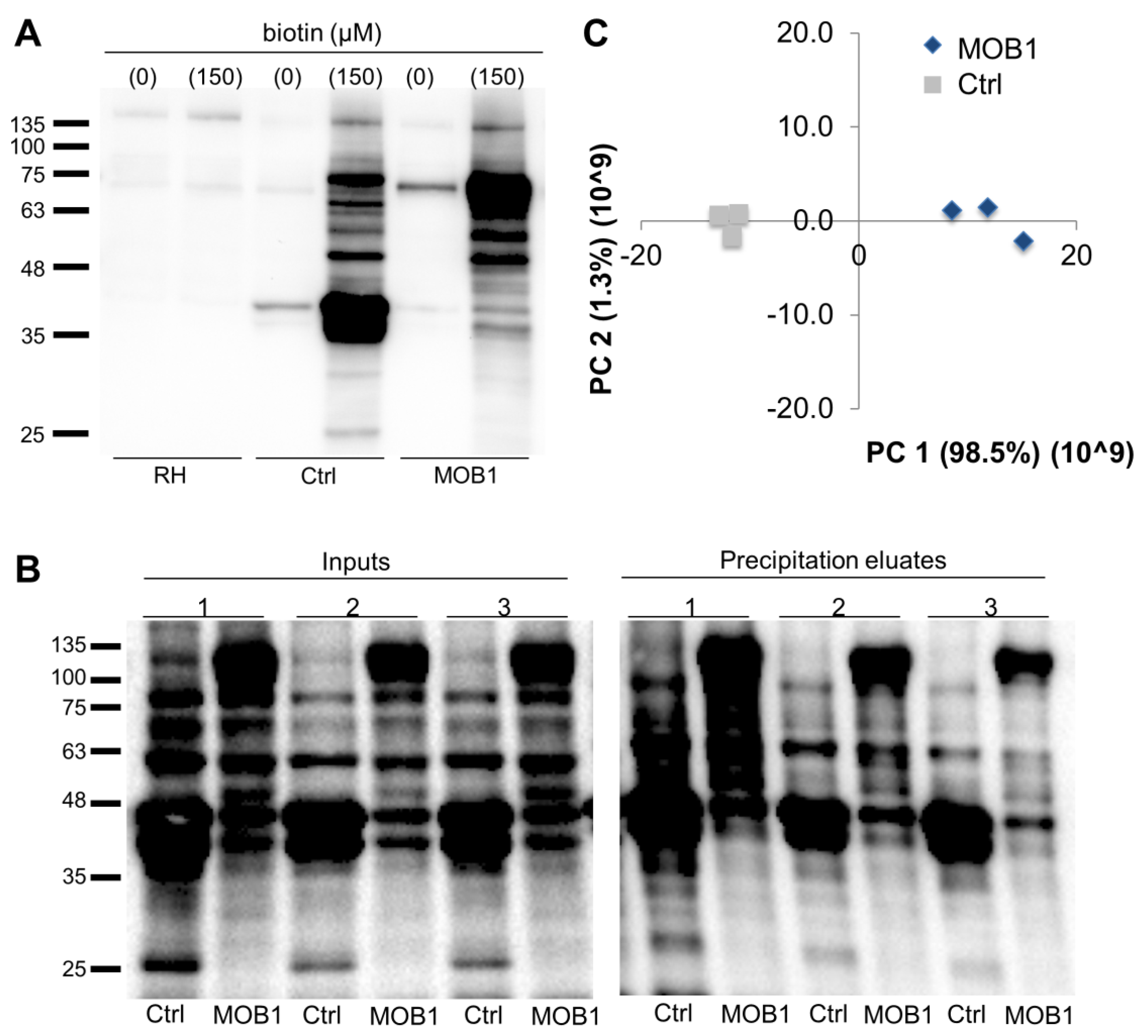



-Western-Blot-NBP2-59938-img0001.jpg)



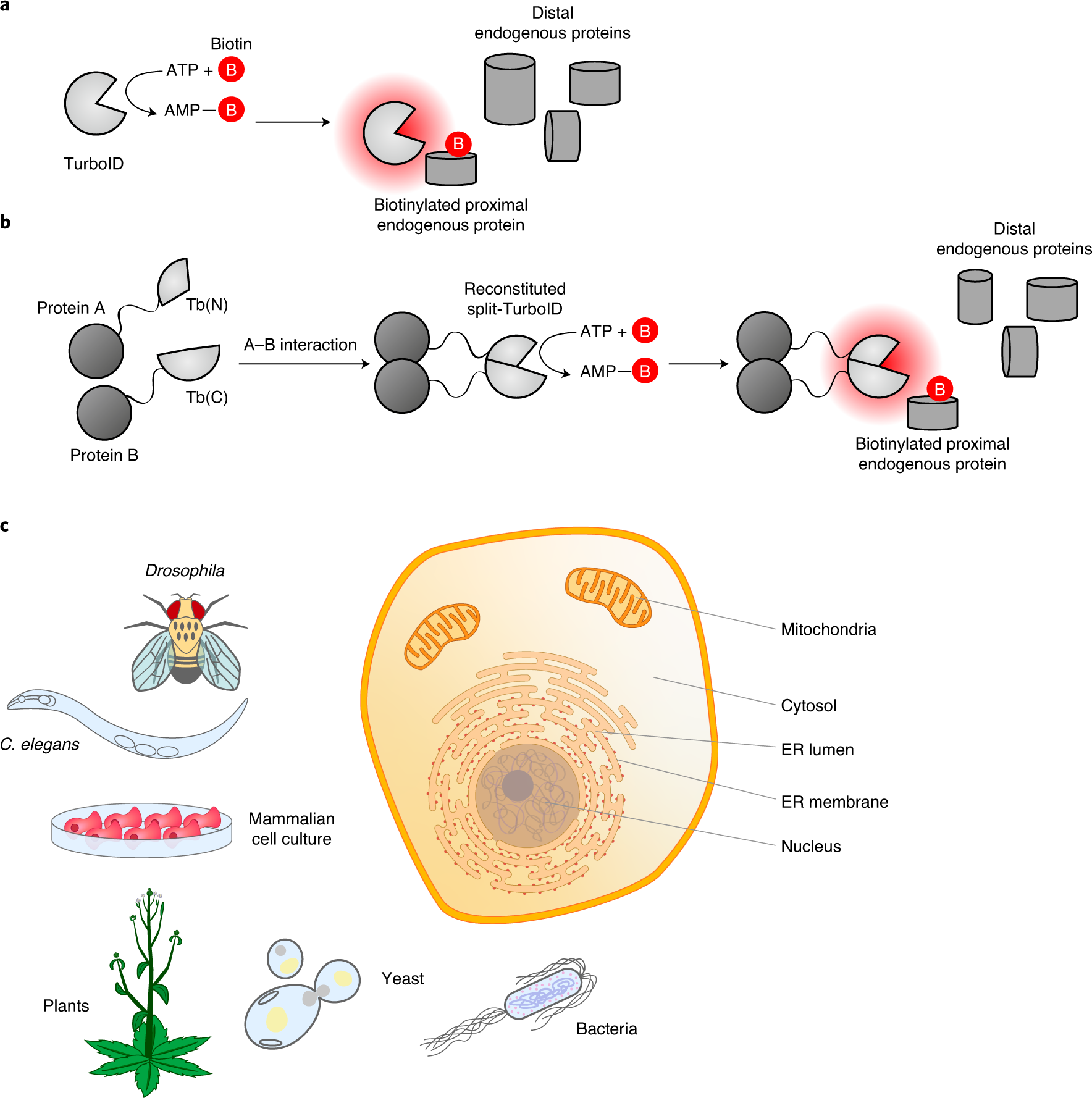


-Western-Blot-NBP2-59939-img0005.jpg)




